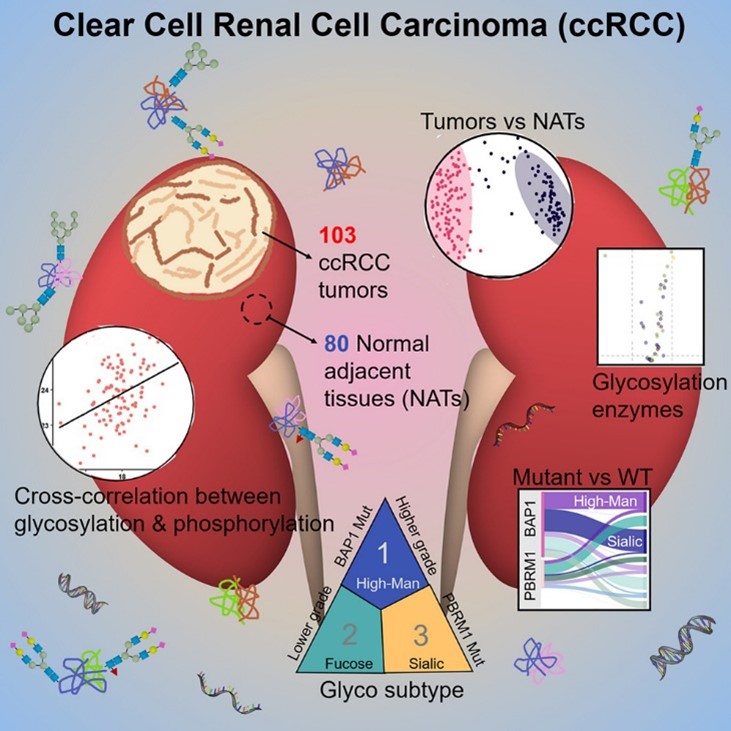
Glycosylation is a ubiquitous type of protein modification that is associated with biological functions and diseases, including cancer. Often found on cell surface or secreted into circulation, glycoproteins can be leveraged for diagnosis and treatment. Recently, a team of researchers from the Clinical Proteomic Tumor Analysis Consortium (CPTAC) conducted a study to understand the role of glycoproteins in the development of clear cell renal cell carcinoma (ccRCC) using patient samples from 103 ccRCC tumors and 80 normal adjacent tissues and resulting in the identification and quantification of over 44,000 intact N-linked glycopeptides from 1,429 glycoproteins (see graphical abstract below). Their remarkable findings, featured in Cell Reports, include key differences in glycosylation patterns between cancerous and normal tissues, variation in glycosylation between high-grade and low-grade ccRCC tumors, different glycosylation in tumors with BAP1 mutation, distinct glycoproteomic-based tumor subtypes, and the cross-correlation between glycan types and phosphoprotein expression.
 Researchers examined four glycan categories according to their monosaccharide composition: oligomannose (High-Man), fucosylated glycans (Fucose), sialylatedglycans (Sialic), and others.. They identified altered glycosylation in ccRCC compared to non-cancerous tissues, with a high proportion of upregulated intact glycopeptides modified by High-Man and Sialic glycans and downregulated glycopeptides modified by Fucose glycans. The differentially expressed intact glycopeptides originated from 317 different glycoproteins, which were associated with clinically relevant pathways including extracellular matrix receptor interaction, cell adhesion, focal adhesion, complement and coagulation cascades, and cholesterol metabolism.
Researchers examined four glycan categories according to their monosaccharide composition: oligomannose (High-Man), fucosylated glycans (Fucose), sialylatedglycans (Sialic), and others.. They identified altered glycosylation in ccRCC compared to non-cancerous tissues, with a high proportion of upregulated intact glycopeptides modified by High-Man and Sialic glycans and downregulated glycopeptides modified by Fucose glycans. The differentially expressed intact glycopeptides originated from 317 different glycoproteins, which were associated with clinically relevant pathways including extracellular matrix receptor interaction, cell adhesion, focal adhesion, complement and coagulation cascades, and cholesterol metabolism.
High-grade tumors were dominated by High-Man glycans, while low-grade tumors were enriched with Sialic glycans. The upregulated High-Man glycans were found on glycoproteins associated with biological processes such as regulation of leukocyte activation, receptor-mediated endocytosis, and integrin-mediated signaling pathway. Sialic glycans were found on glycoproteins associated with cell-substrate adhesion and angiogenesis. To investigate the cross-correlation between glycosylation and phosphorylation in ccRCC, the team classified glycans into four major glycan types and found proteins with co-existing phosphorylation and glycosylation activities, including epidermal growth factor receptor (EGFR), PECAM1, and kidney tissue-associated proteins. Together with other findings, this indicates co-regulation and crosstalk between glycosylation and phosphorylation in ccRCC that has not been described before.
A comparative analysis between BAP1-mutant PBRM1-mutant and wild-type tumors showed specific patterns of glycosylation associated with gene mutations. In BAP1-mutant tumors, elevated glycoproteins were associated with immunity, inflammation, and leukocyte migration. In PBRM1-mutant tumors, elevated glycoproteins were related to metabolic processes. These associations indicate that glycosylation may underlie clinical features, such as aggressiveness, that are unique to specific mutations.
In summary, this glycoproteomic analysis of a large-scale clinical cohort of ccRCC tumors and paired NATs resulting in the identification and quantification of over 44,000 intact N-linked glycopeptides from 1,429 glycoproteins. The study showed that ccRCC tumors exhibited significant differences in glycan type distribution and pathways associated with corresponding glycoproteins compared to NATs. These findings may aid the development of new approaches to stratify patients with ccRCC as well as novel diagnostic and treatment options for ccRCC patients. On the breadth and potential impact of this publication, first author Dr. Mamie Lih wrote, “This study not only demonstrates the role of glycosylation in ccRCC in association with genomic, transcriptomic, proteomic, and phosphoproteomic changes, it also provides an invaluable resource for the ccRCC and broader scientific community for future translational research.”

