RAS gene mutations and their pathways have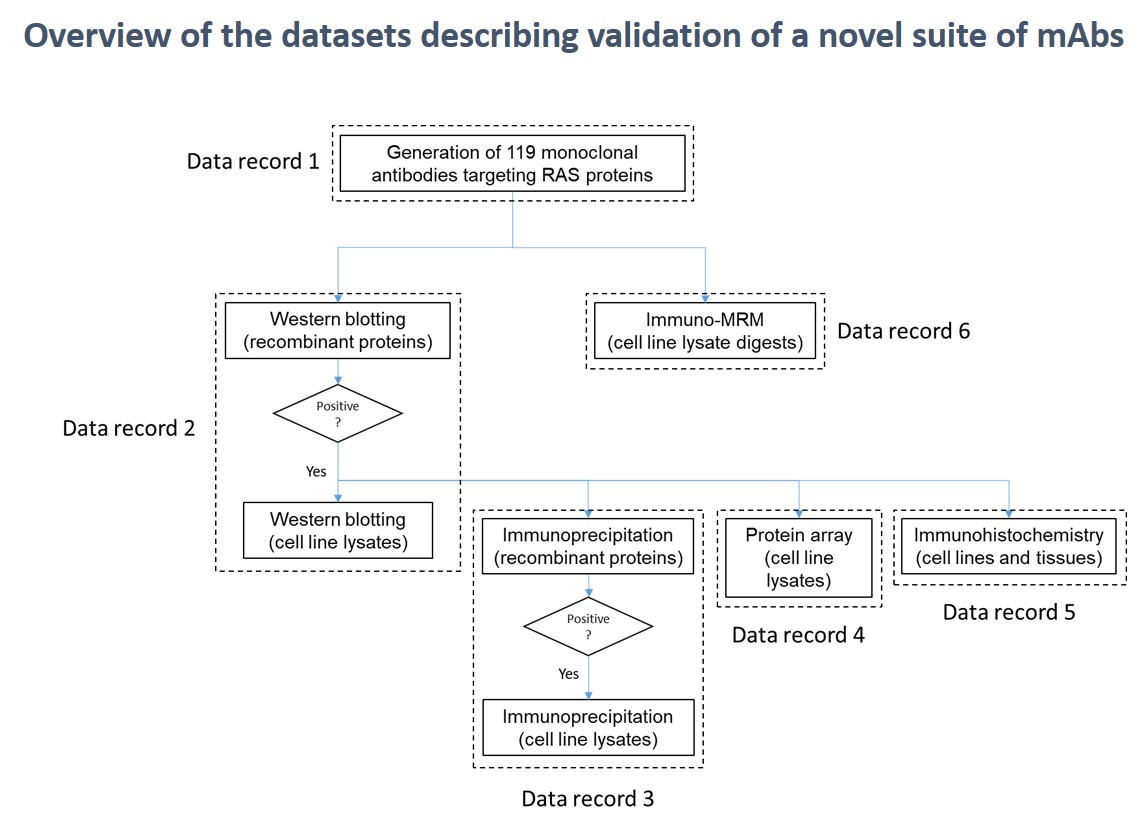 stymied cancer researchers for decades. RAS-driven cancers make up more than 30% of all human cancers due to their association with uncontrollable cell growth, death-signal evasion, and drug resistance. In 2014, the National Cancer Institute launched the RAS Initiative to tackle the befuddling effects of RAS mutations in RAS-driven cancers. The goal of this Initiative is to accelerate scientific discovery for eventual therapeutic use.
stymied cancer researchers for decades. RAS-driven cancers make up more than 30% of all human cancers due to their association with uncontrollable cell growth, death-signal evasion, and drug resistance. In 2014, the National Cancer Institute launched the RAS Initiative to tackle the befuddling effects of RAS mutations in RAS-driven cancers. The goal of this Initiative is to accelerate scientific discovery for eventual therapeutic use.
Joining forces with the RAS Initiative, CPTAC set out to develop immuno-MRM quantitative assays to quantify proteins and phosphoproteins involved in RAS signaling. This signaling pathway normally operates through phosphorylation (and dephosphorylation) of its members in order to activate oncogenesis. To provide a useful resource, assay target selection included the most impactful proteins involved in cancer, selected by RAS Initiative investigators.
In their recent paper published in the journal Scientific Data, part of Nature Publishing Group, CPTAC investigators Amanda Paulovich, Regine Schoenherr and team describe the first step towards assay development: the development of highly characterized monoclonal antibodies. Working with the CPTAC Antibody Program, they developed more than 100 monoclonal antibodies, which were validated by Western blotting, protein immunoprecipitation, protein array analysis, and cell and tissue immunohistochemistry. Additionally, validation was performed in tumor samples, cell lines and/or cancer models. The antibodies were further characterized by their ability to enable detection of endogenous analyte peptides by immuno-MRM mass spectrometry.
Using these antibodies, researchers were able to successfully target 27 phosphopeptides and 69 unmodified peptides from 20 RAS network proteins. This list includes heavy hitters such as AKT, BRAF and ERBB proteins, with many antibodies capable of detecting unmodified and phosphorylated proteoforms. Among them, investigators developed a monoclonal antibody that identified 8 different phosphorylation states of MAPK1. All antibodies will be available to the research community through the Developmental Studies Hybridoma Bank (DSHB) at the University of Iowa. Additionally, all antibody information and characterization data are also publicly available through Panorama Public Repository and/or PRIDE databases.
These antibodies have been developed to be used as well-characterized tools for the cancer research community to study the RAS biological network. A follow-up study detailing the incorporation of these antibodies into immuno-MRM assays with cross-laboratory validation will be available soon. CPTAC is proud to provide these new reagents on our Antibody Portal to advance the discovery of therapies against RAS-driven cancers.

