The CPTAC research group led by Dr. David Fenyö at NYU Langone Medical Center has demonstrated the feasibility of a machine learning image processing tool designed to assist pathologists classifying endometrial cancer. Their customized multi-resolution deep convolutional neural network (CNN) model was able to provide information about patients’ histological subtypes, molecular subtypes, and mutation status rapidly and reliably from digitized H&E-stained pathological images. Just published in Cell Reports Medicine, their results show that CNN models have the potential for clinical application with further refinement to help pathologists determine subtypes and mutations of endometrial carcinoma without the need for sequencing.
Due to the profound 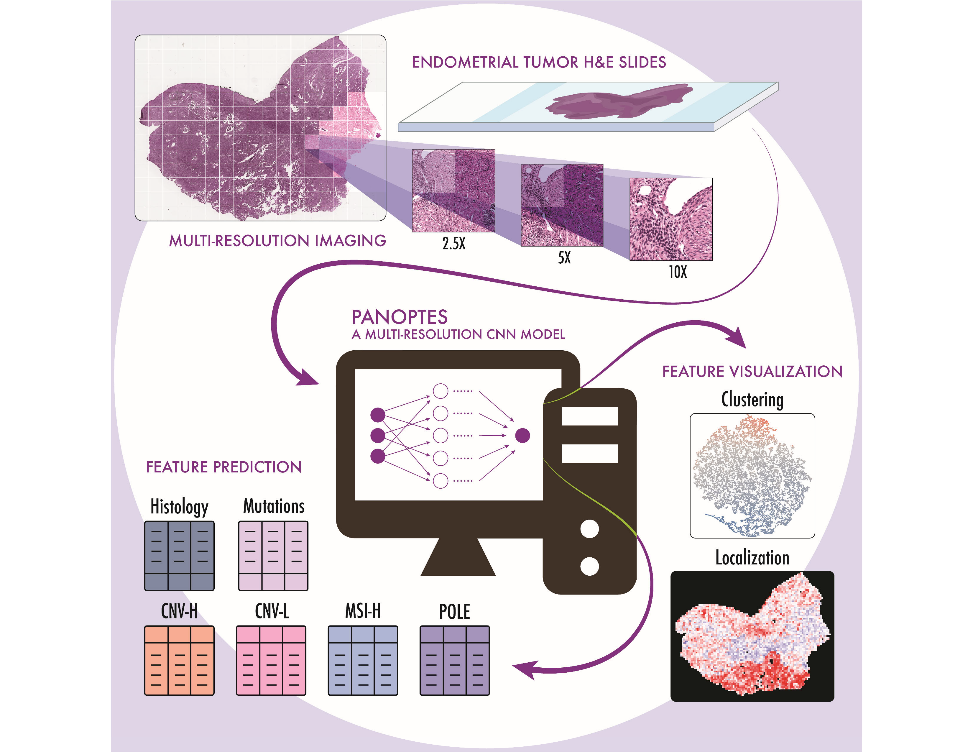 effect certain subtypes of endometrial cancer have on patient prognosis, it is critical to determine tumor subtype for each patient in order to discern the best treatment plan. Currently, examining H&E slides is still the most widely used technique for pathologists to confirm histological subtypes in the clinical setting. Sequencing provides pathologists with additional insights regarding molecular subtypes and mutations, but the availability of this information is limited by the time and cost of this process. Alternatively, computational pathology may provide an alternative, powerful tool which can provide support to pathologists. CNN models have proven capable of segmenting cells in histopathology slides and classifying them into different types based on their morphology—Dr. Fenyö’s group further explored this potential with their customized multi-resolution deep convolutional neural network model, Panoptes.
effect certain subtypes of endometrial cancer have on patient prognosis, it is critical to determine tumor subtype for each patient in order to discern the best treatment plan. Currently, examining H&E slides is still the most widely used technique for pathologists to confirm histological subtypes in the clinical setting. Sequencing provides pathologists with additional insights regarding molecular subtypes and mutations, but the availability of this information is limited by the time and cost of this process. Alternatively, computational pathology may provide an alternative, powerful tool which can provide support to pathologists. CNN models have proven capable of segmenting cells in histopathology slides and classifying them into different types based on their morphology—Dr. Fenyö’s group further explored this potential with their customized multi-resolution deep convolutional neural network model, Panoptes.
Panoptes was able to predict endometrial cancer histological and molecular subtypes as well as mutation status of critical genes with remarkable precision. This performance also generalized well on independent test sets—strong indicator of clinical potential. Classifying endometrioid and serous histological subtypes from datasets of digitized H&E-stained sides resulted in an AUC score of 0.969, and Panoptes can readily distinguish the most lethal molecular subtype, CNV-H with high accuracy (AUC 0.934). Panoptes can also precisely identify CNV-H samples from a histologically endometrioid carcinoma (AUC 0.958), which is widely considered to be one of the most complex patient subgroups in endometrial cancer subtyping. Additionally, Fenyö’s group demonstrated it is possible to reveal and visualize the features which their model learned to classify H&E images through tSNE, a dimensionality reduction technique. These histopathological features were found to be mostly human interpretable, suggesting that they could be incorporated into pathological diagnostic standards and contribute to improved treatment of endometrial carcinoma in the future. Their most optimized models are theoretically capable of facilitating pathologists in real time, taking just under 4 minutes to analyze an H&E slide. By utilizing this iterative approach, the strengths of machines and humans are compounded; Dr. Fenyö believes that “machines and humans can help each other to more effectively learn how to diagnose cancer.”
Next steps for the group include refining the Panoptes architecture to improve the overall performance; the models need to be trained on a more diversified dataset to meet the more stringent criteria for real-world clinical application. They also plan to develop a more advanced Graphical User Interface (GUI) which operates in a fast and user-friendly way such that it can be deployed and tested in a pathologist’s clinical practice. They also have concurrent projects applying Panoptes-based models to predict features in other types of cancers, such as glioblastoma, melanoma, and lung cancer; “Our long-term goal,” says Fenyö, “is to develop a fast, cheap, and accurate method for guiding treatment of different types of cancer.”
Source
Runyu Hong, Wenke Liu, Deborah DeLair, Narges Razavian, David Fenyö. Predicting endometrial cancer subtypes and molecular features from histopathology images using multi-resolution deep learning models. Published online September 21, 2021.

