CPTAC Glioblastoma Multiforme (GBM) Discovery Study Data Release
Proteomic and phosphoproteomic data for the  CPTAC Glioblastoma Discovery Study is now available on the CPTAC Data Portal.
CPTAC Glioblastoma Discovery Study is now available on the CPTAC Data Portal.

COVID-19 is an emerging, rapidly evolving siituation.
What people with cancer should know: https://www.cancer.gov/coronavirus
Guidance for cancer researchers: https://www.cancer.gov/coronavirus-researchers
Get the latest public health information from CDC: https://www.coronavirus.gov
Get the latest research information from NIH: https://www.nih.gov/coronavirus
Proteomic and phosphoproteomic data for the  CPTAC Glioblastoma Discovery Study is now available on the CPTAC Data Portal.
CPTAC Glioblastoma Discovery Study is now available on the CPTAC Data Portal.
And there's more!!! Another new batch of CPTAC histopathology imaging data has been released and is now publicly available on the TCIA CPTAC Pathology Portal.
histopathology imaging data has been released and is now publicly available on the TCIA CPTAC Pathology Portal.
As part of NIH’s mission for collaboration and team science, Dr. Matthew Ellis, Co-Principal Investigator of the CPTAC Proteogenomic Translational Research Center and Dr.
science, Dr. Matthew Ellis, Co-Principal Investigator of the CPTAC Proteogenomic Translational Research Center and Dr.
Did you know about NCI’s Proteomic  (CPTAC) Assay Portal? It’s a free resource for biological scientists performing mass spectroscopy. The portal houses more than 2300 fit-for-purpose, proteomic assays designed and optimized for anyone to use. Each assay is validated using CPTAC experimental g
(CPTAC) Assay Portal? It’s a free resource for biological scientists performing mass spectroscopy. The portal houses more than 2300 fit-for-purpose, proteomic assays designed and optimized for anyone to use. Each assay is validated using CPTAC experimental g
The Proteomic Data Commons (PDC) has released data from the Children's Brain Tumor Tissue Consortium (CBTTC): Pediatric Brain Tumor Atlas dataset!

A new batch of CPTAC histopathology imaging data has been released and is now publicly available on the TCIA CPTAC Pathology Portal.
The TCIA quarterly report lists the addition of 76 new pathology slides and 25 new pathology subjects for the cutaneous melanoma, glioblastoma multiforme, and sarcoma tumor types.
TCIA has partnered with CPTAC to host both the radiology and pathology imaging data generated by the CPTAC Program. Imaging data is available for 10 cancer types including AML, clear cell renal cell carcinoma, head & neck squamous cell carcinoma, lung squamous cell carcinoma, lung adenocarcinoma, ductal carcinoma and corpus endometrial carcinoma. The TCIA CPTAC Pathology Portal has over 5000 histopathology and radiology slides from almost 1500 proteogenomically analyzed tumor cases.
For help navigating the TCIA CPTAC Pathology Portal, please refer to the TCIA CPTAC Pathology Portal Tutorial.
Data Release-20 from the Genomic Data Commons (GDC) includes genomics from a new CPTAC project!
The CPTAC Proteogenomic Confirmatory Study examined prospectively collected colon, breast and ovarian tumors for analysis, including harmonized whole exome sequencing, RNA-Seq, and miRNA-Seq that is now available. Complimentary proteomic and phosphoproteomic analysis can also be found on the CPTAC Data Portal and the Proteomic Data Commons (PDC).
CPTAC has provided the Genomic Data Commons (GDC) with genomic data from over 600 cancer patients with diverse disease types including Lung Adenocarcinoma, Endometrial, Renal, Breast, Colon and Ovarian cancers. The GDC harmonized DNA sequences from CPTAC using GDC DNA-Seq Analysis Pipelines and mRNA Analysis Pipelines. CPTAC proteomic data are processed through the CPTAC Common Data analysis Pipeline (CDAP).
Investigators with the 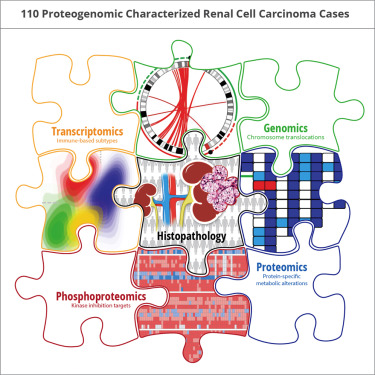 Clinical Proteomic Tumor Analysis Consortium (CPTAC) have generated the most comprehensive multi-omics dataset for clear cell renal cell carcinoma (ccRCC), the most commonly diagnosed kidney cancer subtype. The investigators used integrative proteogenomics, to reveal new insights into kidney cancer.
Clinical Proteomic Tumor Analysis Consortium (CPTAC) have generated the most comprehensive multi-omics dataset for clear cell renal cell carcinoma (ccRCC), the most commonly diagnosed kidney cancer subtype. The investigators used integrative proteogenomics, to reveal new insights into kidney cancer.
Over three hundred proteomic explorers and Clinical Proteomics Tumor Analysis Consortium (CPTAC) participants from around the country convened on the NIH Bethesda campus to share cutting-edge proteogenomic research in the first ever, public-facing CPTAC Scientific Symposium.
Large multi-omics datasets are becoming increasingly 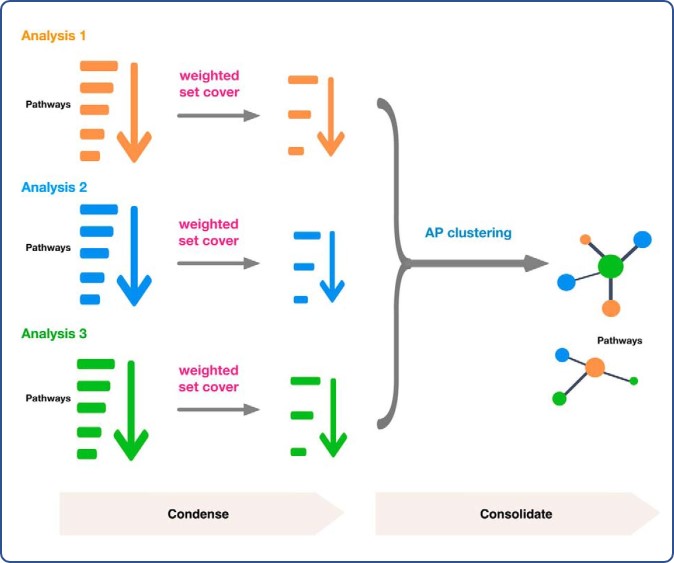 popular for studying biological systems including the identification of biological pathways, or more broadly defined gene sets, that are associated with certain biological or clinical features of interest.
popular for studying biological systems including the identification of biological pathways, or more broadly defined gene sets, that are associated with certain biological or clinical features of interest.